PZLAST-MAG Help
1. Submit a new job.

-
Using PZLAST-MAG, you can search protein sequences against large-scale metagenome-assembled genome (MAG) datasets.
- Query File
- Only a FASTA-formatted file for **protein** sequences is acceptable. (not for DNA sequences) Either a single FASTA or MultiFASTA-formatted file is accepted as input. Paste the input sequence into the box (B) or click the button (C) to select a file.
- Parameters
- PZLAST has the following parameters.
- "Output top N hits" (D)
- This controls how many hits will be output per input sequence. You can set it for top hit (output=1) only, but higher values will give more enjoyable results.
- E-value cut-off (E)
- You can set the cutoff value for the E-value, which defaults to 1e-8. Lowering this value allows you to narrow down the search results to sequences with higher similarity, although it may result in fewer hits.
- Press the submit button (F) to register the job.
- Input file size limit
- All sequences must be >= 10 AA (amino acids) and <= 10,000 AA. The number of input sequences must be <= 10,000. The number of total amino acids must be <= 100,000. The number of possible output must be <= 1,000,000.
Go to the top page or click "Query" in navigation tabs (A).
-
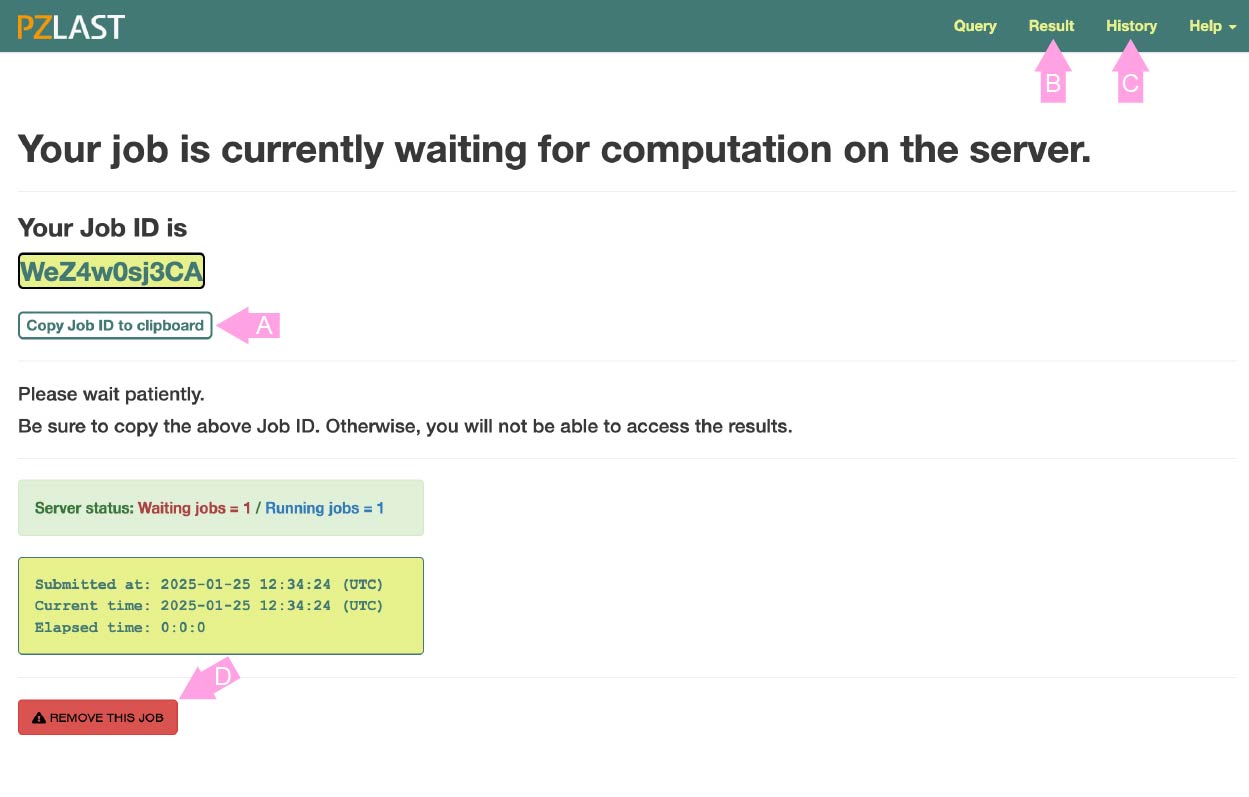
After registering the job, the screen changes to the job status page.
This page shows whether the job is currently WATING to execute the calculation on ZettaScaler-3.0 or is RUNNING on it.
This page is refreshed every few seconds. If the calculation is completed (or unfortunately ends with an error or no-hit), this page will automatically transition to the results page.
All jobs are managed with a unique job ID issued at the time of registration.
If you leave this page, press the copy button (A) and copy the job ID to the clipboard (or somewhere else). You can return to this page again by click the "Result" in the navigation tab (B) and entering the job ID. You can also go back to past jobs from the "History" page (C). Please note that jobs are deleted two weeks after registration.
Note: This site uses cookies to record your job history. Only job ID and registration time are recorded in cookie. If "History" is not displayed, please enable cookies in your browser.
Click the "REMOVE THIS JOB" button (D) if you want to delete the waiting job.
2. Result pages.
- Information page
-
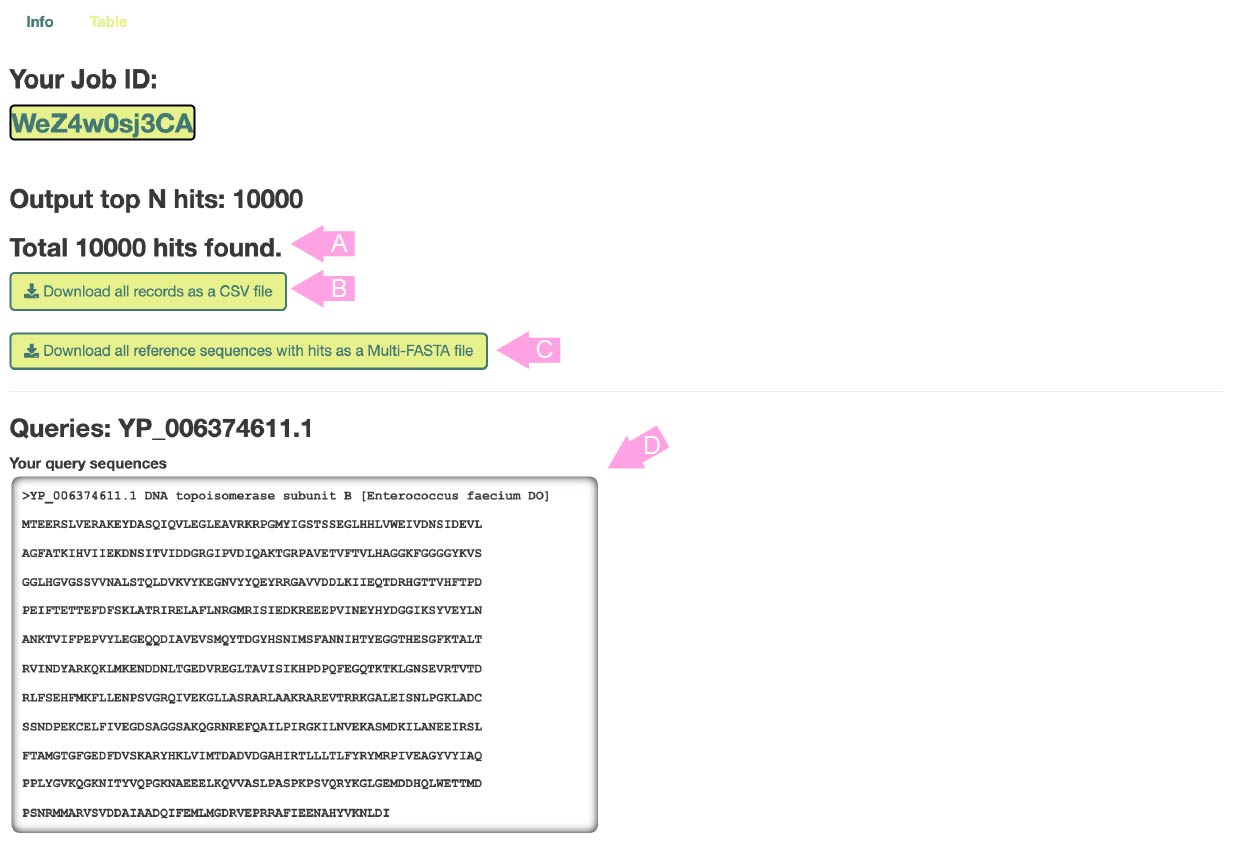
When the calculation is completed, this page will be displayed first.
The total number of hits (or the sum of hits when multi-FASTA is input) is displayed in (A).
Press button (B) to download all results in CSV format.
Press button (C) to download all the hit reference sequences in Multi-FASTA format.
The input information is displayed in (D). All results are distinguished by the sequence ID displayed in "Queries", so remember these IDs when checking the results page below.
- Table results
-
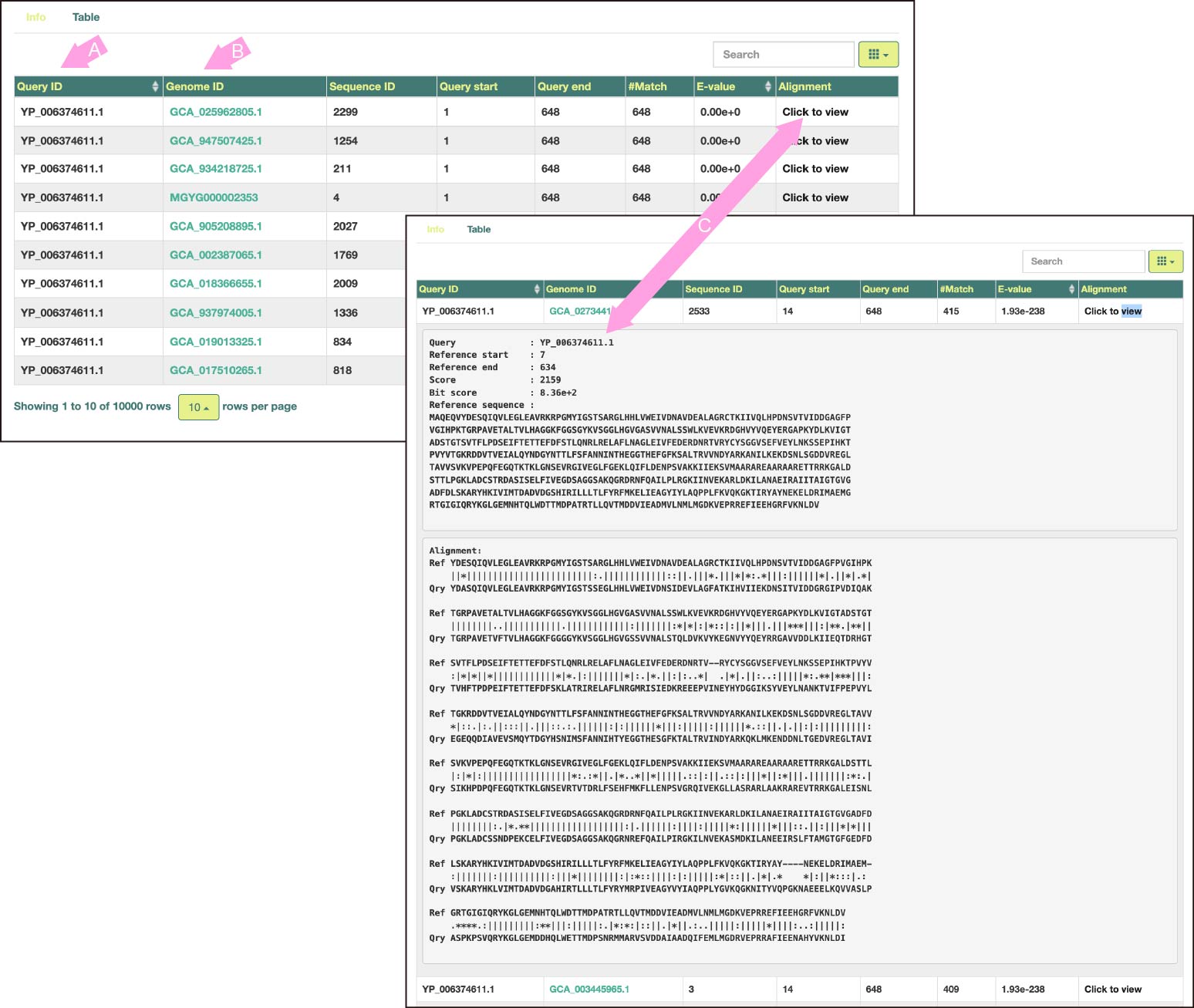
Search results are displayed in a similar tabular format as tools such as BLAST.
By default, hit records are arranged in ascending order of E-value per input sequence.
(A) is the ID of the input sequence. Clicking on the header of this column will sort the table by query names.
(B) is the Genome ID of MAG. Click to jump to the corresponding page of Microbiome Datahub or EMBL-EBI MGnify (depending on which DB the genome data originated from).
(C) For each record, expand the reference protein sequence that the query hit, and the alignment details.
The letters in the alignment mean the following:
'|' ... Match
':' ... BLOSUM62 score > 0
'.' ... BLOSUM62 score = 0
'*' ... BLOSUM62 score < 0
- MEO content
-
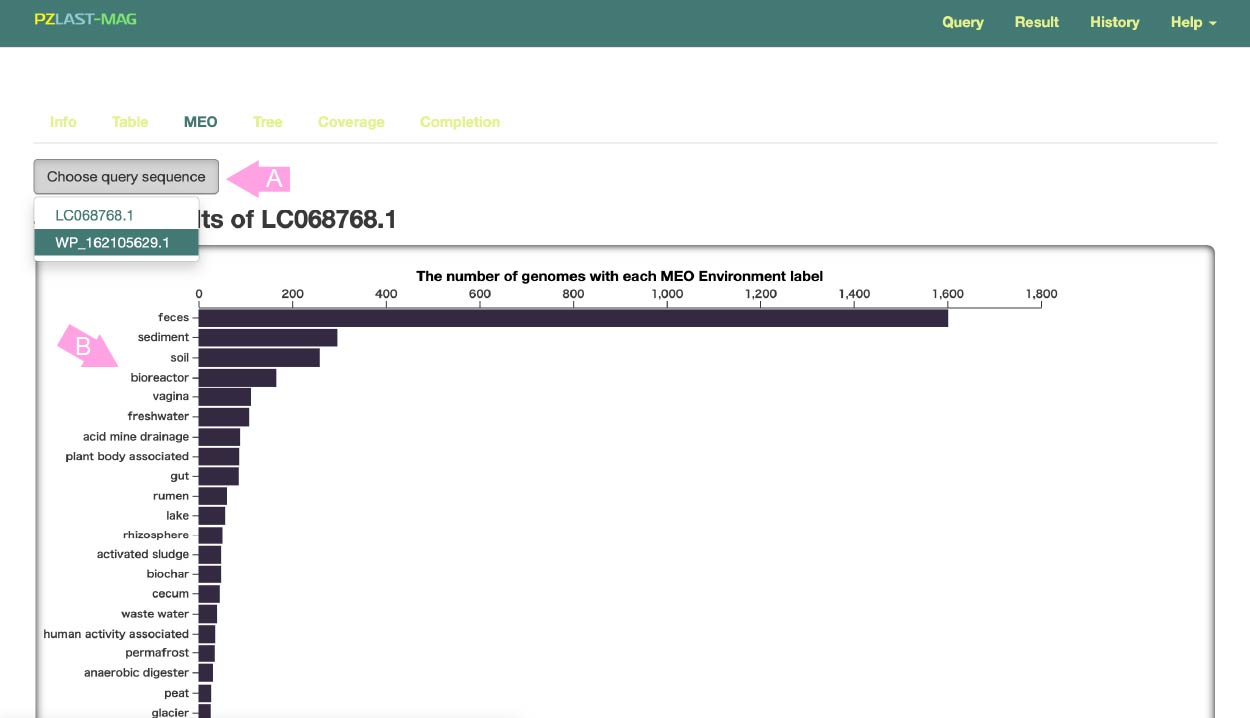
Subsequent pages show the results for **each** query sequence. To switch the query, press the button (A) and select the query for which you want to display the results.
In MicrobiomeDataHub , all metagenomic samples are annotated by MEO . MEO (Metagenome and Microbes Environmental Ontology) is a unified ontology describing what natural or human symbiotic environment the sample was taken from.
This bar chart shows how many MAGs associated with a given MEO were hit by the query (B).
- Phylogenetic tree
-
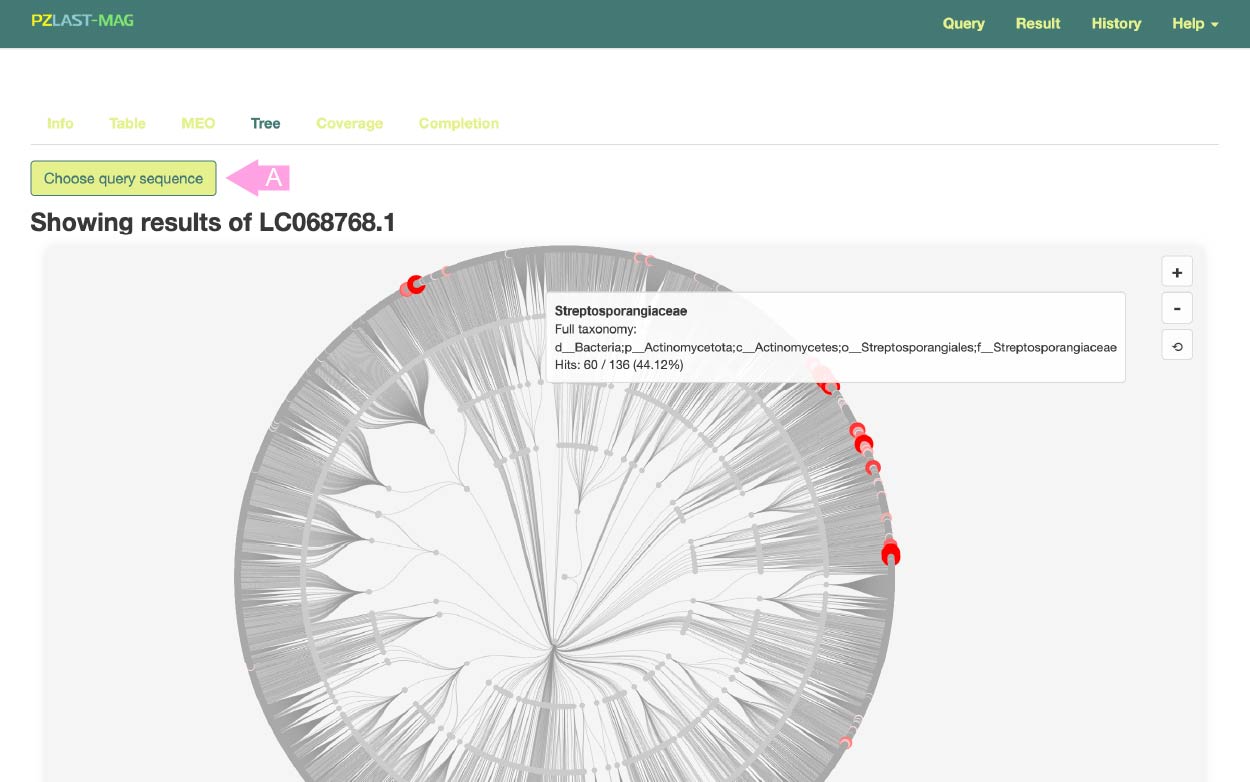
To switch the query, press the button (A) and select the query for which you want to display the results.
This results page visualizes the phylogenetic distribution of hits on an phylogenetic tree.
The evolutionary relationships of Bacteria and Archaea down to the family level are displayed as a tree, with nodes corresponding to hit MAGs highlighted in red. Hovering over a node on the tree displays detailed taxonomic information.
- Coverage
-
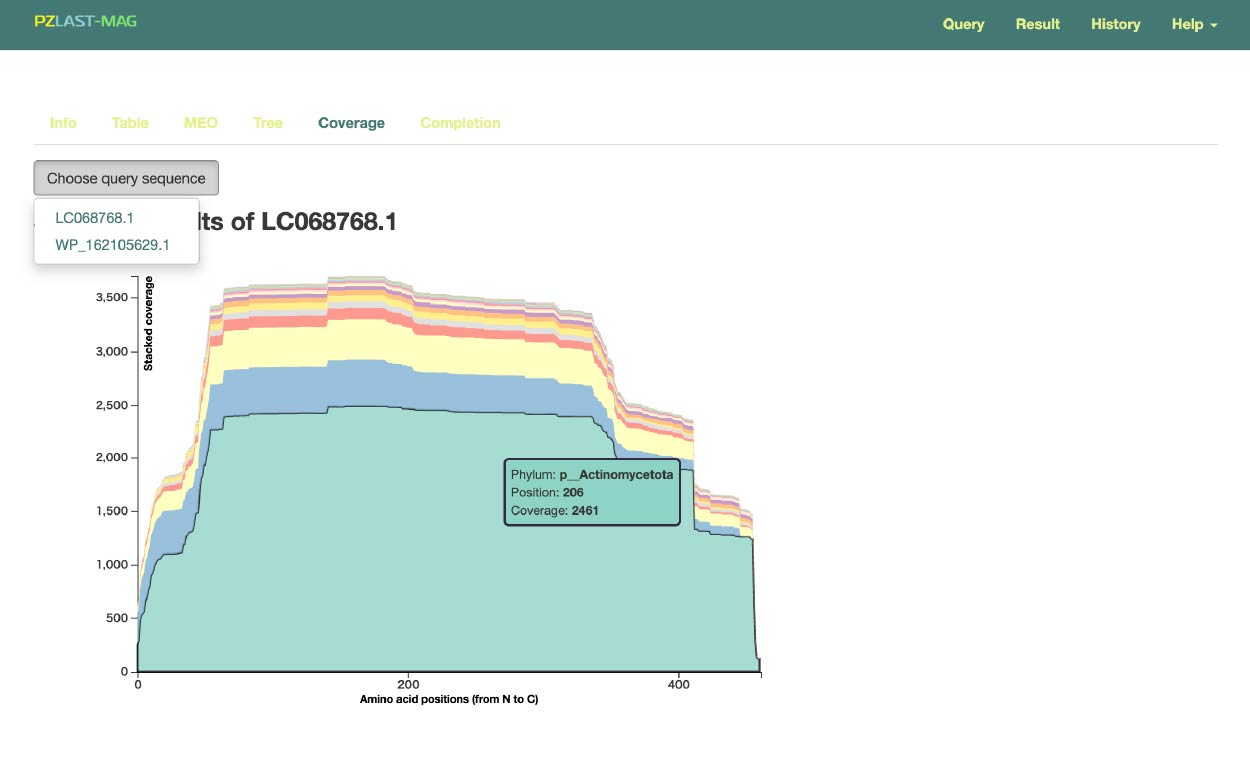
To switch the query, press the button (A) and select the query for which you want to display the results.
This page displays the "taxonomic coverage" for each amino acid position from the N-terminus (left end) to the C-terminus (right end) of the inputted protein query. This may allow for the discovery of relationships where certain domains are found only in specific taxa.
- Completion table
-
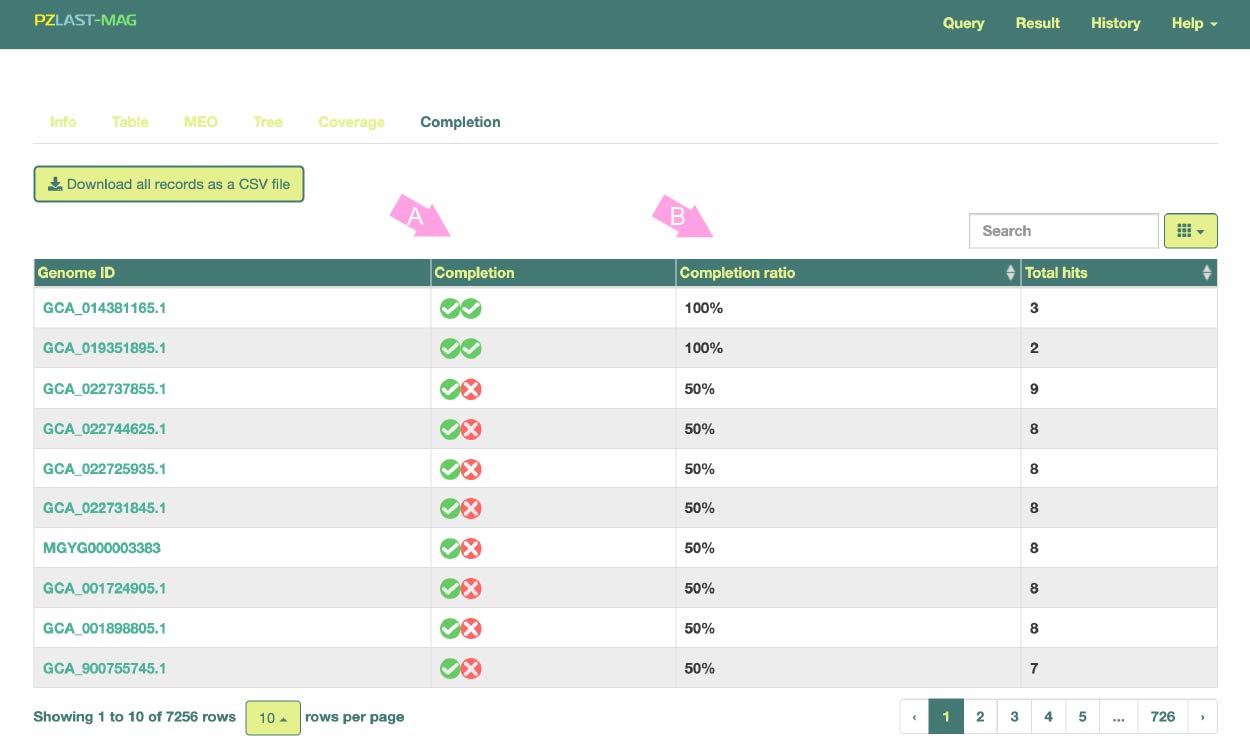
In this page, instead of displaying individual results for hits of the query sequence to specific sequences in a MAG, aggregated results based on the total number of hits per MAG are shown.
Hit MAGs are sorted and displayed based on the "Completion ratio" of multiple protein sequences queried in Multi-FASTA format.
Here, the completion ratio refers to the proportion of the multiple query protein sequences found together in the single MAG, with a MAG considered "found" if at least one sequence hits.
This feature enables, for example, the identification of MAGs in which multiple enzyme protein sequences constituting a certain metabolic pathway are collectively found.
(B) indicates the presence or absence of the query protein in each MAG. (C) represents the Completion ratio of each MAG.